Colloquium
Upcoming Colloquium
Neutrinos from Nuclear Reactors: Status and Outlook
Dr. Juan Pedro Ochoa-Ricoux, UC Irvine
April 29, 2024
11:00am in LA4-120

Abstract
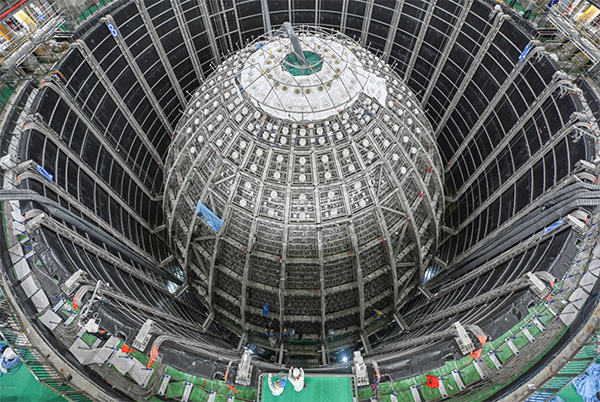
Neutrinos are fascinating elementary particles that help us understand the makeup of our universe as well as the astrophysical and terrestrial sources that produce them. As the most in-tense human-made source of neutrinos ever built, nuclear reactors have been instrumental in studying these elusive particles since their discovery in 1956. In this talk, I will give an overview of the current landscape in reactor neutrino physics. I will begin with a description of neutrinos, the phenomenon of neutrino oscillation, and the most pressing questions in our field. I will then discuss what we have learned from past- and current-generation reactor neutrino experiments, followed by the opportunities offered by a next-generation project currently under construction that is taking the field to the next level in terms of scale and complexity.

The Colloquium is a unique opportunity for students to learn about new developments in physics and what physicists do after they graduate. Hosted by the Physics and Astronomy Department at California State University, Long Beach, the weekly meetings invite guests from universities, research laboratories, and industry to present and discuss current topics in physics. All students are encouraged to attend for a well-rounded experience and training in physics.
Colloquium Coordinator
For information and suggestions about the colloquium please contact the colloquium coordinator:
Dr. Alex Klotz
Alex.Klotz@csulb.edu
Schedule
The following is the schedule for Spring 2024.
| Date | Title | Speaker and Affiliation |
|---|---|---|
| April 29, 2024 | Neutrinos from Nuclear Reactors: Status and Outlook | Dr. Juan Pedro Ochoa-Ricoux, UC Irvine |
| May 6, 2024 | End of Semester Presentations | Students, CSU Long Beach |
Previous Colloquia
| Date | Title | Speaker and Affiliation |
|---|---|---|
| April 22, 2024 | Topological Tuning of DNA-Based Active Matter | Dr. Rae Robertson-Anderson, University of San Diego |
| April 15, 2024 | Gravitational Wave Sources at the Heart of Galaxies | Dr. Smadar Naoz, UCLA |
| April 8, 2024 | Energy: The True Final Frontier (Distinguished Lecture in Physics) | Dr. Ramamoorthy Ramesh, Rice University |
| April 8, 2024 | Vortices, Skyrmions and Cycloids: A New Era in Ferroelectrics | Dr. Ramamoorthy Ramesh, Rice University |
| March 27, 2024 | Gravitational Waves from f-modes as a Tool to Probe the Neutron Star Interior | Dr. Debarati Chatterjee, Inter-University Centre for Astronomy and Astrophysics |
| March 25, 2024 | Exploring Topological Phase Transitions and Dynamic Strain Engineering in Quantum Materials | Dr. Luis A. Jauregui, UC Irvine |
| March 18, 2024 | DNA Liquids | Dr. Omar Saleh, UC Santa Barbara |
| March 11, 2024 | The Gravity Tunnel in a Non-Uniform Earth | Dr. Alex Klotz, CSU Long Beach |
| February 26, 2024 | A City on Mars: Can We Settle Space, Should We Settle Space, and Have We Really Thought This Through? | Zach Weinersmith, illustrator and writer for Saturday Morning Breakfast Cereal |
| February 19, 2024 | Driving Quantum Matter to Extremes | Dr. Sarah Grefe, CSU Long Beach |
| February 12, 2024 | Solar System Archaelogy: Multi-disciplinary Approaches to Understanding Planetary System Formation and Environmental Science | Dr. Gerardo Dominguez, CSU San Marcos |
| February 5, 2024 | Assembly, Disassembly, and Mechanics of Entropic Colloidosomes | Dr. Zvonimir Dogic, UC Santa Barbara |
| January 29, 2024 | What Quantum Materials Can Reveal When Interrogated with Photoemission and Electronic Transport Probes | Dr. Claudia Ojeda-Aristizaba, CSU Long Beach |
The Colloquium Archive has the Colloquia from previous semesters.
Sponsors
We acknowledge with gratitude donations and support from the following present sponsors:
- H.E. and H.B. Miller and Family Endowment
- Benjamin Carter
- American Physical Society
- Anonymous
We also acknowledge with gratitude our past donors: The Forty-Niner Shops, Inc., The Northrop Grumman Foundation, Sandra Dana, Anonymous.
If you wish to support the Colloquium, please contact the colloquium coordinator or the department chair. Thank you!